Support
Synchronizing Image Sets Before Segmentation
Important: Image syncing must be completed before performing segmentation.
Important: Image syncing must be completed before performing segmentation.
1. Open the Image Tabs
Open the Controller Menu.
Select the Images tab.
Under both the Primary and Secondary image tabs, you’ll see a clock icon.
Clicking this icon opens the Phase Timing Settings menu, where you can set parameters to synchronize both image sets.
2. Set the Cardiac Cycle Start Point (Primary Image)
In the Phase Timing Settings, look for the green arrow above the phase box.
This arrow marks the reference point that defines the start of the cardiac cycle.
Use the left and right toggles next to Cycle Start Index to adjust the start of the cardiac phase.
For this example, set the beginning of the cardiac cycle to atrial systole.
3. Adjust the Secondary Image Timing
Open the Phase Time Settings for the Secondary Image.
In an echocardiography (echo) series, we want to display one full cardiac cycle—the opening and closing of the valve.
Cycle through the image phases to identify the start and end of one cardiac cycle.
Use Trim from Start or Trim from End to remove any extra phases outside of the single cycle.
In this example, trimming is not necessary.
4. Synchronize with the CT Scan
Compare the cardiac phase of the CT scan with that of the echo.
In the CT scan, the heart is in atrial systole, where the mitral valve is open.
Open the Phase Timing Settings for the Secondary Image.
Using the Cycle Start Index, locate the image phase showing the open mitral valve.
You’ll see the green arrow move over this phase — indicating synchronization with the CT scan.
5. Verify Synchronization
Visually inspect both image sets to confirm the phases align.
Adjust the Cycle Start Index for the secondary image if needed.
6. Proceed to Segmentation
Once both image sets are synchronized, you can begin segmenting your model.
Separating Models into Individual Structures and Colours
The Select/Split function in Elucis is a powerful and efficient method for separating complex models into individual anatomical structures.
This will guide you through selecting, refining, and splitting portions of a model, as well as transferring selections between structures.
The Select/Split function in Elucis is a powerful and efficient method for separating complex models into individual anatomical structures.
This will guide you through selecting, refining, and splitting portions of a model, as well as transferring selections between structures.
1. Prepare the Model
Open the Controller Menu.
Ensure the model you intend to split is the active structure.
Open the Stylus Menu and press the Select button.
You can now begin selecting the anatomy you wish to isolate — for this example, we’ll focus on the left ventricle.
2. Understand the Preview and Selection Toggles
On the surface plane, you’ll see three toggles:
Show Selection – Displays both the full model and your selection.
Preview Inside – Shows only your selection, hiding the rest of the model.
Preview Outside – Shows the model minus your selection.
These views, combined with the Grow and Shrink Selection tools, allow for highly precise control.
3. Refine the Selection
While in Preview Inside mode:
You’ll see your selection only.
Use Grow Selection to add anatomy to your selection — visible anatomy will appear as it’s added.
Use Shrink Selection to remove anatomy from your selection.
While in Preview Outside mode:
You’ll see the model without your current selection.
Use Grow Selection to remove anatomy from the model (adding it to the selection).
Use Shrink Selection to add anatomy back to the model (removing it from the selection).
These complementary tools enable accurate boundary definition between two adjacent structures.
4. Split the Model
Once satisfied with your selection:
Turn to the Work Surface.
You’ll see two buttons: Split and Copy.
Next to the Split button, locate the Destination toggle.
Ensure the setting below it reads New Structure.
Press Split.
Your selection will change color, indicating it has been separated into a new structure.
To verify:
Open the Models Menu — you’ll see a new structure labeled “Intersection.”
Rename the new structure as desired.
Repeat these steps if you wish to segment your model further.
5. Transfer a Selection Between Structures
To transfer part of one structure into another:
Ensure the source structure (the one containing the anatomy you want to move) is active.
Use the Select Tool to highlight the portion you wish to transfer.
On the Work Surface, locate the Destination toggle next to the Split button.
Cycle through the available structures until you find the target structure.
Press Split.
The selected anatomy will now transfer to the chosen destination structure.
This process provides precise, flexible control over model organization, allowing for cleaner segmentation and easier anatomical analysis.
Segmenting a 4D Heart Model
Follow these steps to create and segment a 4D heart model from your image series.
Follow these steps to create and segment a 4D heart model from your image series.
1. Create a New Structure
Open the Models tab in the Controller Menu.
Create a new structure.
2. Access the Image Series
Open the Images tab.
Scroll to the bottom to locate the Time Series Settings.
This section allows you to:
Set the image series to Static (manually cycle through phases), or
Set it to Dynamic (automatically cycle through phases at adjustable speeds).
To segment a 4D heart, you’ll need to segment all phases in the image series.
For efficiency, set the series speed to its maximum before beginning.
3. Begin Segmenting
Open the Stylus Menu and select the Add Tool.
On the surface plane, ensure Use Threshold and Display Threshold are both enabled.
Adjust the threshold to match the contrast-filled heart chambers.
Begin segmenting — you’ll see the segmentation applied across all image phases.
Continue cycling through the image volume while segmenting until all desired anatomy is built.
Tip: Segmentation can also be performed in 3D, though this can be computationally demanding and may cause temporary performance lag.
4. Clean Up the Segmentation
Once the 4D blood pool is segmented, you can clean up floating anatomy for a smoother model.
Open the Images tab and set Time Series Settings to Static.
Open the Stylus Menu and select the Select-Connected Tool.
While squeezing the index finger trigger on your dominant hand, tap the blood volume you want to select.
A white film will appear around the model, indicating the selected blood volume.
When the selection is complete, tap Delete Unselected on the surface plane.
This removes all floating anatomy and yields a tidier blood volume.
Repeat this process for each phase.
5. Build a Shell Around the Blood Volume
Open the Operations tab in the Controller Menu.
Under Single Structure Operations:
Select the blood volume as the structure.
Change the operation to Shell.
Set your desired shell thickness.
Before executing, ensure Apply to All Phases is enabled.
Press Execute.
A new structure will appear, and you’ll see it listed in the Models tab.
Screen Capture and Recording
Screen capture and recording in Elucis allow you to document and export specific model views for use in presentations, educational materials, or quick visual references.
This guide covers how to record videos, take snapshots, and capture rotational image series from both the desktop and VR environments.
Screen capture and recording in Elucis allow you to document and export specific model views for use in presentations, educational materials, or quick visual references.
This guide covers how to record videos, take snapshots, and capture rotational image series from both the desktop and VR environments.
1. Navigate and Prepare Your Model
Before capturing:
Rotate the model using the right mouse button.
Zoom in/out using the scroll wheel.
Center the model by pressing both the scroll wheel and right mouse button simultaneously.
2. Adjust Screen Visualization
On the right-hand side of the screen, open the Screen Visualization Options tab.
Here, you can:
Show or hide the surface plane and image volume box.
Adjust and orient the light source.
3. Access the Content Recording Tools
Open the Content Recording tab to view the three available capture modes:
Start Video Recording
Screen Snapshot
Rotational Series
4. Record a Video
Press Start Video Recording.
A flashing “Recording” signal will appear at the top of the screen to indicate active recording.
Rotate and manipulate the model as needed using the mouse.
When finished, open the Content Recording tab and press Stop Video Recording.
A small zoom visualization near the Snapshots and Recordings button confirms the video has been saved to the project.
5. Capture a Still Image
Open the Content Recording tab.
Press Screen Snapshot.
The same zoom visualization confirms that the capture has been saved.
6. Review and Export Captures
Open the Snapshots and Recordings button on the left-hand side of the screen.
Click any thumbnail to open a review window where you can inspect and add annotations.
To export:
Click the top-left circle on each thumbnail you want to export, or press Select All.
Click Export Selected and choose a save location on your PC.
All captures and recordings remain stored within the project and can be accessed later.
7. Capture a Snapshot in VR
Open the Stylus Menu and press the Camera button.
A shutter screen will appear — grab and position it as desired.
On the surface plane, you can:
Adjust the field of view.
Show or hide the surface plane and image volume box.
Switch the background between white and black.
To capture the image, squeeze the index finger trigger on your dominant hand.
A shutter sound confirms the capture.
To review and export, return to the desktop and open the Snapshots and Recordings menu.
8. Capture a Rotational Series
Open the Content Recording tab and select Rotational Series.
Choose the export format — either PC image or DICOM files.
DICOM files can also be exported directly to an integrated PACS system.
Configure the rotational capture parameters:
Field of view
Centering
File format
Image resolution
Number of rotational steps
Series name (optional)
Preview the capture using the Preview button.
When satisfied, press Export to save the series to your PC or PACS system.
Rendering and Positioning an Echo Image Set
Follow these steps to volume render an echo image set and accurately position it within a segmented heart model.
Follow these steps to volume render an echo image set and accurately position it within a segmented heart model.
1. Open the Echo Image Set
Open the Controller Menu.
Select the Images tab.
Locate the image tab that contains the Echo image set and press Volume Rendering.
Open the drop-down menu and select the Echo preset.
2. Adjust the Rendered Model
You can refine the detail and appearance of the rendered model using the following controls:
Crop sliders: Adjust the visible portion of the volume.
Offset sliders: Modify the depth and position of the rendering.
3. Apply a Mask to Shape the Cardiac Valve
To refine the valve’s contours and shape:
Open the Stylus Menu.
At the bottom, find the Mask tool section.
Press Add Mask to begin shaping the rendered model.
Use the middle finger trigger on your non-dominant hand to toggle between Add Mask and Remove Mask modes.
Continue refining until the valve’s appearance is satisfactory.
4. Reposition the Valve Within the Segmented Model
Open the Controller Menu again.
Go to the Images tab and select Image Registration.
Enable the Use Free Hand toggle.
This allows you to freely manipulate and position the rendered valve within the segmented heart model.
5. Final Positioning and Alignment
Open the Images tab and disable the Clippable toggle.
Create a new clip plane that intersects the segmented heart model.
Use this plane to position the valve precisely.
For improved accuracy, align the model in relation to the CT scan using the planar surface as a reference.
Thresholding Tutorial
Setting a proper threshold is one of the most important steps when segmenting a model that accurately reflects the anatomy of interest. The Threshold Scroll Bar in Elucis allows you to define values that correspond to the specific tissue densities you wish to include in your segmentation.
Setting a proper threshold is one of the most important steps when segmenting a model that accurately reflects the anatomy of interest. The Threshold Scroll Bar in Elucis allows you to define values that correspond to the specific tissue densities you wish to include in your segmentation.
1. Preparing to Set a Threshold
Open the Controller Menu.
Click Models, then create a New Structure.
On the Stylus Menu, select the Add button.
At the bottom left of the Surface Plane, select Display Threshold.
You’ll now see regions highlighted in pink — these areas represent your current threshold range, which can be modified using the scroll bar in the center of the surface plane.
2. Understanding the Threshold Scroll Bar
The scroll bar has two slider buttons — a left and a right — that define the minimum and maximum range of your threshold:
Moving the sliders closer together creates a narrow threshold range, capturing a smaller density window.
Moving them further apart creates a wider threshold range, encompassing more densities.
3. General Threshold Ranges (CT Data)
When working with CT image sets, the scroll bar roughly corresponds to tissue density in Hounsfield units (HU):
Left side → Radiolucent structures (air, fat, low density).
Middle → Soft tissue (muscle, organs).
Right side → Radiodense tissue (bone, contrast media, metal).
Example:
If segmenting the myocardium while excluding the contrast within heart chambers:
Move the left slider until the myocardium appears highlighted.
Move the right slider until the contrast regions are no longer highlighted.
The threshold range will narrow, indicating that only the densities within that range are included.
4. Thresholding in MR and Echo Data
For MRI and Echocardiography datasets:
Use the left slider to broadly capture the tissue of interest.
Adjust the right slider to refine and isolate the desired region.
This process allows flexible adaptation to different intensity scales used in non-CT modalities.
5. Fine-Tuning Threshold Values
For greater precision:
Gently drag the slider buttons for small adjustments, or
Use the Minimum and Maximum toggles located on the right-hand side of the surface plane.
These fine controls allow you to capture subtle tissue boundaries accurately.
6. Reducing Noise
If the threshold appears noisy or contains unwanted speckles:
Check the Reduce Noise box.
You’ll see that the threshold becomes smoother, with cleaner, more defined borders.
Note that noise reduction applies only to the threshold visualization, not the image data itself.
By mastering thresholding, you ensure that your segmentations closely match true anatomical boundaries — providing a clean, accurate foundation for further refinement and analysis.
Aligning Multiple Image Sets
When multiple image sets are imported, Elucis will automatically align them using their DICOM information.
However, in some cases, the images may not align automatically. Follow the steps below to manually align them.
When multiple image sets are imported, Elucis will automatically align them using their DICOM information.
However, in some cases, the images may not align automatically. Follow the steps below to manually align them.
1. Automatically Align Image Sets
If the images are not aligned after import:
Open the Images tab.
Select Image Registration.
On the surface plane, locate Registration Type and select Automatic.
Press the Auto Register button.
This will automatically align the secondary image to the primary image.
2. Copy Alignment to Additional Image Sets
If you have multiple secondary image sets, you can copy the alignment from the first secondary image and apply it to the others.
Open the Images tab.
From the Secondary Image Set dropdown, select the series you want to align.
On the surface plane, locate Copy Registration From, followed by the name of the previously aligned secondary image set.
Press Copy to align the image to the same position as the previous set.
Repeat this process for all remaining secondary image sets.
Manual Image Co-Registration
When both primary and secondary image sets are imported, Elucis automatically detects their shared DICOM properties and aligns the images accordingly.
In cases where images do not align automatically—or when manual adjustment is desired to highlight specific anatomy—you can use the Image Registration tools.
When both primary and secondary image sets are imported, Elucis automatically detects their shared DICOM properties and aligns the images accordingly.
In cases where images do not align automatically—or when manual adjustment is desired to highlight specific anatomy—you can use the Image Registration tools.
1. Access the Image Registration Tools
Open the Controller Menu on your non-dominant hand.
Select the Images tab.
You’ll see both your Primary and Secondary image sets, along with Fusion Settings at the bottom of the menu.
Fusion Settings let you:
Display the Primary, Secondary, or a blend of both image sets.
Use different blending modes to enhance anatomical visibility.
2. Open the Image Registration Panel
In the Images tab, select Image Registration.
On the surface plane, you’ll find two registration options for aligning the images.
Note: Adjustments only affect the secondary image set. The primary image remains fixed.
3. Use the Movement Gizmo
The Movement Gizmo provides three key tools for manual alignment:
Translation Tool: Move the secondary image along the X, Y, and Z axes.
Rotation Tool: Rotate the image around the same axes.
Spin Tool: Fine-tune the image orientation.
These tools allow precise manipulation to achieve perfect alignment.
4. Fine-Tune Image Positioning
For even greater accuracy, use the precision tools available on the surface plane:
Extra Precise Mode: Requires larger stylus movements for small positional changes.
1 mm / 1° Mode: Enables movement in exact 1-millimeter or 1-degree increments.
5. Align Image Sets in 3D
To align in three dimensions:
In the Images tab, enable Ghost View for both the Primary and Secondary image sets.
Use the Movement Gizmo to line up visible anatomical borders on the ghosted views.
Once alignment looks correct, press Save State on the surface plane.
This action locks both image sets into position.
Importing DICOM Images from Your PC
Follow these steps to import DICOM images into your project:
1. Create a New Project
Click New Project.
Name your project.
2. Select Image Location
Press Select Path.
Navigate through your PC to the folder where your DICOM images are stored.
3. Review Your Images
Once the images have uploaded, use the scroll bar on the right to cycle through them.
Confirm that the image set is correct.
4. Finalize Import
Press Create to complete the import.
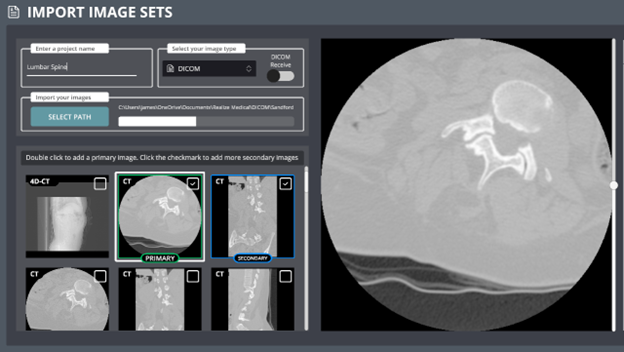
Managing Multiple DICOM Image Sets
If you’ve imported multiple DICOM images:
Set a Primary Image
Double-click on an image thumbnail to make it your primary image.
A green box will appear around the thumbnail.Set Secondary Images
Check the box in the upper-right corner of a thumbnail to mark it as secondary.
A blue box will appear around the thumbnail.
You can add as many secondary image sets as you need.